Find us on Google Scholar, ResearcherID, and MyNCBI.
Complete Publication List
269 results
| [269] | RNA folding and the origins of catalytic activity in the hairpin ribozyme. (Wilson, T. J., Nahas, M., Araki, L., Harusawa, S., Ha, T. and Lilley, D. M.), In Blood Cells Mol. Dis., . [bibtex] [pdf] |
| 2020 | |
| [268] | Crystal structure and ligand-induced folding of the SAM/SAH riboswitch. (Huang, L., Liao, T. W., Wang, J., Ha, T. and Lilley, D. M. J.), In Nucleic Acids Res., 2020. [bibtex] [pdf] |
| [267] | Light-controlled twister ribozyme with single-molecule detection resolves RNA function in time and space. (Korman, A., Sun, H., Hua, B., Yang, H., Capilato, J. N., Paul, R., Panja, S., Ha, T., Greenberg, M. M. and Woodson, S. A.), In Proc. Natl. Acad. Sci. U.S.A., 2020. [bibtex] [pdf] |
| [266] | E. coli Rep helicase and RecA recombinase unwind G4 DNA and are important for resistance to G4-stabilizing ligands. (Paul, T., Voter, A. F., Cueny, R. R., Gavrilov, M., Ha, T., Keck, J. L. and Myong, S.), In Nucleic Acids Res., 2020. [bibtex] [pdf] |
| [265] | ORCA/LRWD1 Regulates Homologous Recombination at ALT-Telomeres by Modulating Heterochromatin Organization. (Hsu, R. Y. C., Lin, Y. C., Redon, C., Sun, Q., Singh, D. K., Wang, Y., Aggarwal, V., Mitra, J., Matur, A., Moriarity, B., Ha, T., Aladjem, M. I., Prasanth, K. V. and Prasanth, S. G.), In iScience, 2020. [bibtex] [pdf] |
| [264] | Just Took a DNA Test, Turns Out 100% Not That Phase. (Gitler, A. D., Shorter, J., Ha, T. and Myong, S.), In Mol. Cell, 2020. [bibtex] [pdf] |
| [263] | Single-Molecule Analysis and Engineering of DNA Motors. (Mohapatra, S., Lin, C. T., Feng, X. A., Basu, A. and Ha, T.), In Chem. Rev., 2020. [bibtex] [pdf] |
| [262] | Very fast CRISPR on demand. (Liu, Y., Zou, R. S., He, S., Nihongaki, Y., Li, X., Razavi, S., Wu, B. and Ha, T.), In Science, 2020. [bibtex] [pdf] |
| 2019 | |
| [261] | Single molecule analysis of effects of non-canonical guide RNAs and specificity-enhancing mutations on Cas9-induced DNA unwinding. (Okafor, I. C., Singh, D., Wang, Y., Jung, M., Wang, H., Mallon, J., Bailey, S., Lee, J. K. and Ha, T.), In Nucleic Acids Res., 2019. [bibtex] [pdf] |
| [260] | Streamlining effects of extra telomeric repeat on telomeric DNA folding revealed by fluorescence-force spectroscopy. (Mitra, J. and Ha, T.), In Nucleic Acids Res., 2019. [bibtex] [pdf] |
| [259] | Structural basis for DNA unwinding at forked dsDNA by two coordinating Pif1 helicases. (Su, N., Byrd, A. K., Bharath, S. R., Yang, O., Jia, Y., Tang, X., Ha, T., Raney, K. D. and Song, H.), In Nat Commun, 2019. [bibtex] [pdf] |
| [258] | Determinants of target prioritization and regulatory hierarchy for the bacterial small RNA SgrS. (Bobrovskyy, M., Azam, M. S., Frandsen, J. K., Zhang, J., Poddar, A., Ma, X., Henkin, T. M., Ha, T. and Vanderpool, C. K.), In Mol. Microbiol., 2019. [bibtex] [pdf] |
| [257] | Nanomechanics and co-transcriptional folding of Spinach and Mango. (Mitra, J. and Ha, T.), In Nat Commun, 2019. [bibtex] [pdf] |
| [256] | Strategy for Compositional Analysis of the Hair Cell Mechanotransduction Complex Using TIRF Microscopy. (Clark, S., Elferich, J., Gai, J., Goehring, A., Mitra, J., Ha, T. and Gouaux, E.), In Microsc. Microanal., 2019. [bibtex] [pdf] |
| [255] | Fight against background noise in stimulated emission depletion nanoscopy. (Ma, Y. and Ha, T.), In Phys Biol, 2019. [bibtex] [pdf] |
| [254] | Extreme mechanical diversity of human telomeric DNA revealed by fluorescence-force spectroscopy. (Mitra, J., Makurath, M. A., Ngo, T. T. M., Troitskaia, A., Chemla, Y. R. and Ha, T.), In Proc. Natl. Acad. Sci. U.S.A., 2019. [bibtex] [pdf] |
| [253] | Nuclear speckle fusion via long-range directional motion regulates speckle morphology after transcriptional inhibition. (Kim, J., Han, K. Y., Khanna, N., Ha, T. and Belmont, A. S.), In J. Cell. Sci., 2019. [bibtex] [pdf] |
| [252] | Junction resolving enzymes use multivalency to keep the Holliday junction dynamic. (Zhou, R., Yang, O., Déclais, A. C., Jin, H., Gwon, G. H., Freeman, A. D. J., Cho, Y., Lilley, D. M. J. and Ha, T.), In Nat. Chem. Biol., 2019. [bibtex] [pdf] |
| [251] | Functional instability allows access to DNA in longer transcription Activator-Like effector (TALE) arrays. (Geiger-Schuller, K., Mitra, J., Ha, T. and Barrick, D.), In Elife, 2019. [bibtex] [pdf] |
| [250] | Hexameric helicase G40P unwinds DNA in single base pair steps. (Schlierf, M., Wang, G., Chen, X. S. and Ha, T.), In Elife, 2019. [bibtex] [pdf] |
| [249] | Structural Mechanisms of Cooperative DNA Binding by Bacterial Single-Stranded DNA-Binding Proteins. (Dubiel, K., Myers, A. R., Kozlov, A. G., Yang, O., Zhang, J., Ha, T., Lohman, T. M. and Keck, J. L.), In J. Mol. Biol., 2019. [bibtex] [pdf] |
| [248] | Contractility kits promote assembly of the mechanoresponsive cytoskeletal network. (Kothari, P., Srivastava, V., Aggarwal, V., Tchernyshyov, I., Van Eyk, J. E., Ha, T. and Robinson, D. N.), In J. Cell. Sci., 2019. [bibtex] [pdf] |
| 2018 | |
| [247] | Publisher Correction: Precision and accuracy of single-molecule FRET measurements-a multi-laboratory benchmark study. (Hellenkamp, B., Schmid, S., Doroshenko, O., Opanasyuk, O., Kühnemuth, R., Adariani, S. R., Ambrose, B., Aznauryan, M., Barth, A., Birkedal, V., Bowen, M. E., Chen, H., Cordes, T., Eilert, T., Fijen, C., Gebhardt, C., Götz, M., Gouridis, G., Gratton, E., Ha, T., Hao, P., Hanke, C. A., Hartmann, A., Hendrix, J., Hildebrandt, L. L., Hirschfeld, V., Hohlbein, J., Hua, B., Hübner, C. G., Kallis, E., Kapanidis, A. N., Kim, J. Y., Krainer, G., Lamb, D. C., Lee, N. K., Lemke, E. A., Levesque, B., Levitus, M., McCann, J. J., Naredi-Rainer, N., Nettels, D., Ngo, T., Qiu, R., Robb, N. C., Röcker, C., Sanabria, H., Schlierf, M., Schröder, T., Schuler, B., Seidel, H., Streit, L., Thurn, J., Tinnefeld, P., Tyagi, S., Vandenberk, N., Vera, A. M., Weninger, K. R., Wünsch, B., Yanez-Orozco, I. S., Michaelis, J., Seidel, C. A. M., Craggs, T. D. and Hugel, T.), In Nat. Methods, 2018. [bibtex] [pdf] |
| [246] | Precision and accuracy of single-molecule FRET measurements-a multi-laboratory benchmark study. (Hellenkamp, B., Schmid, S., Doroshenko, O., Opanasyuk, O., Kühnemuth, R., Rezaei Adariani, S., Ambrose, B., Aznauryan, M., Barth, A., Birkedal, V., Bowen, M. E., Chen, H., Cordes, T., Eilert, T., Fijen, C., Gebhardt, C., Götz, M., Gouridis, G., Gratton, E., Ha, T., Hao, P., Hanke, C. A., Hartmann, A., Hendrix, J., Hildebrandt, L. L., Hirschfeld, V., Hohlbein, J., Hua, B., Hübner, C. G., Kallis, E., Kapanidis, A. N., Kim, J. Y., Krainer, G., Lamb, D. C., Lee, N. K., Lemke, E. A., Levesque, B., Levitus, M., McCann, J. J., Naredi-Rainer, N., Nettels, D., Ngo, T., Qiu, R., Robb, N. C., Röcker, C., Sanabria, H., Schlierf, M., Schröder, T., Schuler, B., Seidel, H., Streit, L., Thurn, J., Tinnefeld, P., Tyagi, S., Vandenberk, N., Vera, A. M., Weninger, K. R., Wünsch, B., Yanez-Orozco, I. S., Michaelis, J., Seidel, C. A. M., Craggs, T. D. and Hugel, T.), In Nat. Methods, 2018. [bibtex] [pdf] |
| [245] | Mimicking Co-Transcriptional RNA Folding Using a Superhelicase. (Hua, B., Panja, S., Wang, Y., Woodson, S. A. and Ha, T.), In J. Am. Chem. Soc., 2018. [bibtex] [pdf] |
| [244] | Toward Single-Cell Single-Molecule Pull-Down. (Wang, X., Park, S., Zeng, L., Jain, A. and Ha, T.), In Biophys. J., 2018. [bibtex] [pdf] |
| [243] | Cdc42-dependent modulation of rigidity sensing and cell spreading in tumor repopulating cells. (Chowdhury, F., Doğanay, S., Leslie, B. J., Singh, R., Amar, K., Talluri, B., Park, S., Wang, N. and Ha, T.), In Biochem. Biophys. Res. Commun., 2018. [bibtex] [pdf] |
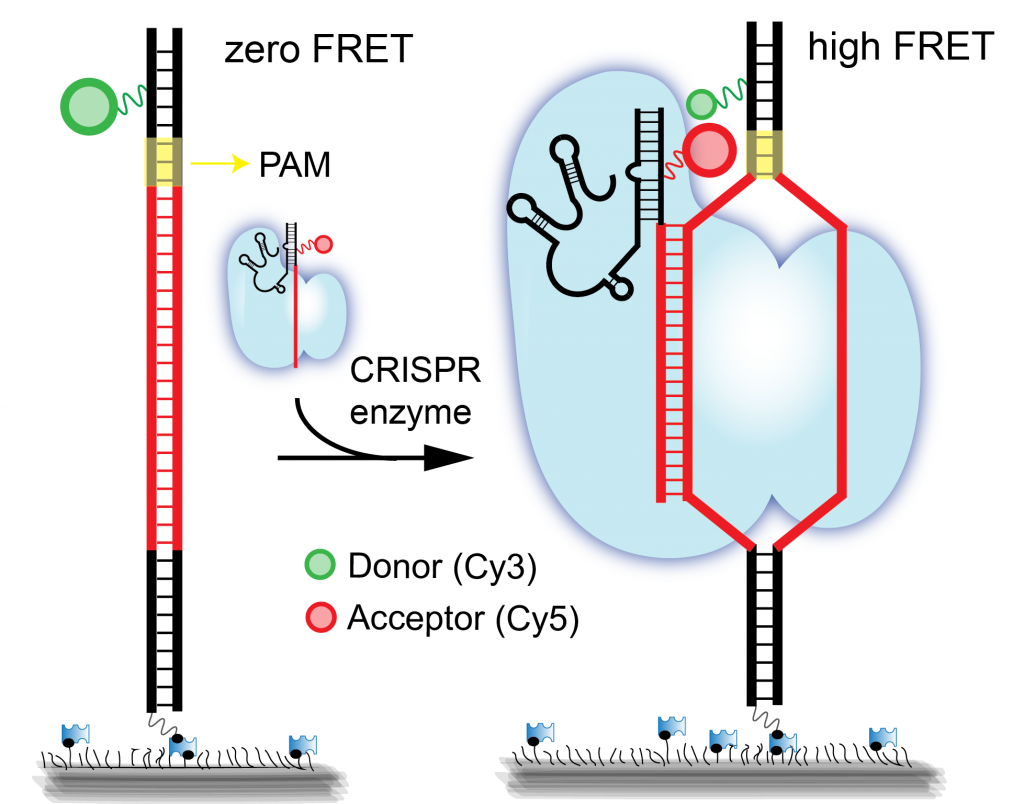
| [242] | Real-time observation of DNA target interrogation and product release by the RNA-guided endonuclease CRISPR Cpf1 (Cas12a). (Singh, D., Mallon, J., Poddar, A., Wang, Y., Tippana, R., Yang, O., Bailey, S. and Ha, T.), In Proc. Natl. Acad. Sci. U.S.A., 2018. [bibtex] [pdf] |
| [241] | Correlating Transcription Initiation and Conformational Changes by a Single-Subunit RNA Polymerase with Near Base-Pair Resolution. (Koh, H. R., Roy, R., Sorokina, M., Tang, G. Q., Nandakumar, D., Patel, S. S. and Ha, T.), In Mol. Cell, 2018. [bibtex] [pdf] |
| [240] | Mechanisms of improved specificity of engineered Cas9s revealed by single-molecule FRET analysis. (Singh, D., Wang, Y., Mallon, J., Yang, O., Fei, J., Poddar, A., Ceylan, D., Bailey, S. and Ha, T.), In Nat. Struct. Mol. Biol., 2018. [bibtex] [pdf] |
| [239] | Understanding the Molecular Mechanisms of the CRISPR Toolbox Using Single Molecule Approaches. (Singh, D. and Ha, T.), In ACS Chem. Biol., 2018. [bibtex] [pdf] |
| [238] | The Single-Molecule Centroid Localization Algorithm Improves the Accuracy of Fluorescence Binding Assays. (Hua, B., Wang, Y., Park, S., Han, K. Y., Singh, D., Kim, J. H., Cheng, W. and Ha, T.), In Biochemistry, 2018. [bibtex] [pdf] |
| [237] | Editorial Overview: Single-Molecule Approaches up to Difficult Challenges in Folding and Dynamics. (Zhang, Y., Ha, T. and Marqusee, S.), In J. Mol. Biol., 2018. [bibtex] [pdf] |
| [236] | TGFβ1 reinforces arterial aging in the vascular smooth muscle cell through a long-range regulation of the cytoskeletal stiffness. (Zhu, W., Kim, B. C., Wang, M., Huang, J., Isak, A., Bexiga, N. M., Monticone, R., Ha, T., Lakatta, E. G. and An, S. S.), In Sci Rep, 2018. [bibtex] [pdf] |
| [235] | Mechanism of polypurine tract primer generation by HIV-1 reverse transcriptase. (Figiel, M., Krepl, M., Park, S., Poznański, J., Skowronek, K., Gołąb, A., Ha, T., Šponer, J. and Nowotny, M.), In J. Biol. Chem., 2018. [bibtex] [pdf] |
| [234] | Single-Molecule Studies of ssDNA-Binding Proteins Exchange. (Yang, O. and Ha, T.), In Meth. Enzymol., 2018. [bibtex] [pdf] |
| [233] | Single-Molecule FRET Analysis of Replicative Helicases. (Lee, S. J., Syed, S. and Ha, T.), In Methods Mol. Biol., 2018. [bibtex] [pdf] |
| 2017 | |
| [232] | Quantitative analysis of multilayer organization of proteins and RNA in nuclear speckles at super resolution. (Fei, J., Jadaliha, M., Harmon, T. S., Li, I. T. S., Hua, B., Hao, Q., Holehouse, A. S., Reyer, M., Sun, Q., Freier, S. M., Pappu, R. V., Prasanth, K. V. and Ha, T.), In J. Cell. Sci., 2017. [bibtex] [pdf] |
| [231] | Single-cell analysis of early antiviral gene expression reveals a determinant of stochastic IFNB1 expression. (Doğanay, S., Lee, M. Y., Baum, A., Peh, J., Hwang, S. Y., Yoo, J. Y., Hergenrother, P. J., García-Sastre, A., Myong, S. and Ha, T.), In Integr Biol (Camb), 2017. [bibtex] [pdf] |
| [230] | Sphingolipids facilitate age asymmetry of membrane proteins in dividing yeast cells. (Singh, P., Ramachandran, S. K., Zhu, J., Kim, B. C., Biswas, D., Ha, T., Iglesias, P. A. and Li, R.), In Mol. Biol. Cell, 2017. [bibtex] [pdf] |
| [229] | Metals induce transient folding and activation of the twister ribozyme. (Panja, S., Hua, B., Zegarra, D., Ha, T. and Woodson, S. A.), In Nat. Chem. Biol., 2017. [bibtex] [pdf] |
| [228] | Single-molecule imaging reveals the translocation and DNA looping dynamics of hepatitis C virus NS3 helicase. (Lin, C. T., Tritschler, F., Lee, K. S., Gu, M., Rice, C. M. and Ha, T.), In Protein Sci., 2017. [bibtex] [pdf] |
| [227] | Voltage-gated sodium channels assemble and gate as dimers. (Clatot, J., Hoshi, M., Wan, X., Liu, H., Jain, A., Shinlapawittayatorn, K., Marionneau, C., Ficker, E., Ha, T. and Deschênes, I.), In Nat Commun, 2017. [bibtex] [pdf] |
| [226] | Evolution of protein-coupled RNA dynamics during hierarchical assembly of ribosomal complexes. (Abeysirigunawardena, S. C., Kim, H., Lai, J., Ragunathan, K., Rappé, M. C., Luthey-Schulten, Z., Ha, T. and Woodson, S. A.), In Nat Commun, 2017. [bibtex] [pdf] |
| [225] | A genetically encoded fluorescent tRNA is active in live-cell protein synthesis. (Masuda, I., Igarashi, T., Sakaguchi, R., Nitharwal, R. G., Takase, R., Han, K. Y., Leslie, B. J., Liu, C., Gamper, H., Ha, T., Sanyal, S. and Hou, Y. M.), In Nucleic Acids Res., 2017. [bibtex] [pdf] |
| [224] | Notch-Jagged complex structure implicates a catch bond in tuning ligand sensitivity. (Luca, V. C., Kim, B. C., Ge, C., Kakuda, S., Wu, D., Roein-Peikar, M., Haltiwanger, R. S., Zhu, C., Ha, T. and Garcia, K. C.), In Science, 2017. [bibtex] [pdf] |
| [223] | Mapping cell surface adhesion by rotation tracking and adhesion footprinting. (Li, I. T., Ha, T. and Chemla, Y. R.), In Sci Rep, 2017. [bibtex] [pdf] |
| [222] | Flipping nanoscopy on its head. (Xiao, J. and Ha, T.), In Science, 2017. [bibtex] [pdf] |
| [221] | Structural biology: Growth factor rattled out of its cage. (Ha, T.), In Nature, 2017. [bibtex] [pdf] |
| [220] | Probing Single Helicase Dynamics on Long Nucleic Acids Through Fluorescence-Force Measurement. (Lin, C. T. and Ha, T.), In Methods Mol. Biol., 2017. [bibtex] [pdf] |
| 2016 | |
| [219] | Single-Molecule Analysis of Lipid-Protein Interactions in Crude Cell Lysates. (Arauz, E., Aggarwal, V., Jain, A., Ha, T. and Chen, J.), In Anal. Chem., 2016. [bibtex] [pdf] |
| [218] | Natural antisense RNA promotes 3' end processing and maturation of MALAT1 lncRNA. (Zong, X., Nakagawa, S., Freier, S. M., Fei, J., Ha, T., Prasanth, S. G. and Prasanth, K. V.), In Nucleic Acids Res., 2016. [bibtex] [pdf] |
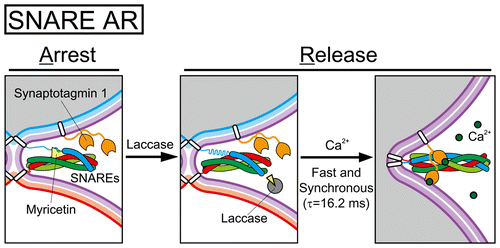
| [217] | A Chemical Controller of SNARE-Driven Membrane Fusion That Primes Vesicles for Ca(2+)-Triggered Millisecond Exocytosis. (Heo, P., Yang, Y., Han, K. Y., Kong, B., Shin, J. H., Jung, Y., Jeong, C., Shin, J., Shin, Y. K., Ha, T. and Kweon, D. H.), In J. Am. Chem. Soc., 2016. [bibtex] [pdf] |
| [216] | Rupture force of cell adhesion ligand tethers modulates biological activities of a cell-laden hydrogel. (Lee, M. K., Park, J., Wang, X., Roein-Peikar, M., Ko, E., Qin, E., Lee, J., Ha, T. and Kong, H.), In Chem. Commun. (Camb.), 2016. [bibtex] [pdf] |
| [215] | Direct evidence for sequence-dependent attraction between double-stranded DNA controlled by methylation. (Yoo, J., Kim, H., Aksimentiev, A. and Ha, T.), In Nat Commun, 2016. [bibtex] [pdf] |
| [214] | Plantazolicin is an ultra-narrow spectrum antibiotic that targets the Bacillus anthracis membrane. (Molohon, K. J., Blair, P. M., Park, S., Doroghazi, J. R., Maxson, T., Hershfield, J. R., Flatt, K. M., Schroeder, N. E., Ha, T. and Mitchell, D. A.), In ACS Infect Dis, 2016. [bibtex] [pdf] |
| [213] | Spider Silk Peptide Is a Compact, Linear Nanospring Ideal for Intracellular Tension Sensing. (Brenner, M. D., Zhou, R., Conway, D. E., Lanzano, L., Gratton, E., Schwartz, M. A. and Ha, T.), In Nano Lett., 2016. [bibtex] [pdf] |
| [212] | Probing Nature's Nanomachines One Molecule at a Time. (Ha, T.), In Biophys. J., 2016. [bibtex] [pdf] |
| [211] | Effects of cytosine modifications on DNA flexibility and nucleosome mechanical stability. (Ngo, T. T., Yoo, J., Dai, Q., Zhang, Q., He, C., Aksimentiev, A. and Ha, T.), In Nat Commun, 2016. [bibtex] [pdf] |
| [210] | The light side of the force. (Basu, A. and Ha, T.), In Elife, 2016. [bibtex] [pdf] |
| [209] | Constructing modular and universal single molecule tension sensor using protein G to study mechano-sensitive receptors. (Wang, X., Rahil, Z., Li, I. T., Chowdhury, F., Leckband, D. E., Chemla, Y. R. and Ha, T.), In Sci Rep, 2016. [bibtex] [pdf] |
| [208] | Structural dynamics of potassium-channel gating revealed by single-molecule FRET. (Wang, S., Vafabakhsh, R., Borschel, W. F., Ha, T. and Nichols, C. G.), In Nat. Struct. Mol. Biol., 2016. [bibtex] [pdf] |
| [207] | Single-molecule fluorescence microscopy of native macromolecular complexes. (Aggarwal, V. and Ha, T.), In Curr. Opin. Struct. Biol., 2016. [bibtex] [pdf] |
| [206] | Real-time observation of DNA recognition and rejection by the RNA-guided endonuclease Cas9. (Singh, D., Sternberg, S. H., Fei, J., Doudna, J. A. and Ha, T.), In Nat Commun, 2016. [bibtex] [pdf] |
| [205] | Correction: Nanoscale mechanics guides cellular decision making. (Rahil, Z., Pedron, S., Wang, X., Ha, T., Harley, B. and Leckband, D.), In Integr Biol (Camb), 2016. [bibtex] [pdf] |
| [204] | Nanoscale mechanics guides cellular decision making. (Rahil, Z., Pedron, S., Wang, X., Ha, T., Harley, B. and Leckband, D.), In Integr Biol (Camb), 2016. [bibtex] [pdf] |
| [203] | In Planta Single-Molecule Pull-Down Reveals Tetrameric Stoichiometry of HD-ZIPIII:LITTLE ZIPPER Complexes. (Husbands, A. Y., Aggarwal, V., Ha, T. and Timmermans, M. C.), In Plant Cell, 2016. [bibtex] [pdf] |
| [202] | Ebola Virus Does Not Induce Stress Granule Formation during Infection and Sequesters Stress Granule Proteins within Viral Inclusions. (Nelson, E. V., Schmidt, K. M., Deflubé, L. R., Doğanay, S., Banadyga, L., Olejnik, J., Hume, A. J., Ryabchikova, E., Ebihara, H., Kedersha, N., Ha, T. and Mühlberger, E.), In J. Virol., 2016. [bibtex] [pdf] |
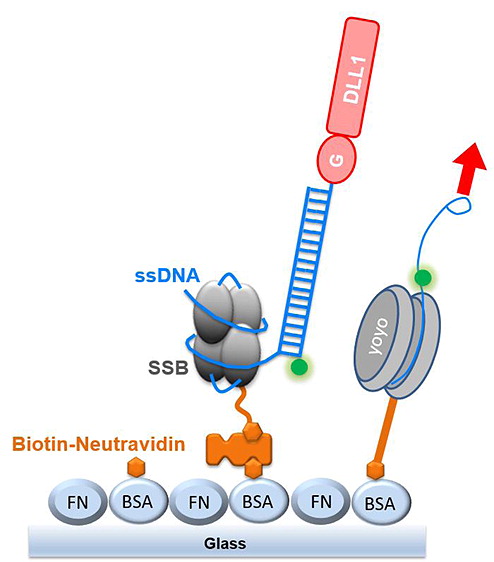
| [201] | Defining Single Molecular Forces Required for Notch Activation Using Nano Yoyo. (Chowdhury, F., Li, I. T., Ngo, T. T., Leslie, B. J., Kim, B. C., Sokoloski, J. E., Weiland, E., Wang, X., Chemla, Y. R., Lohman, T. M. and Ha, T.), In Nano Lett., 2016. [bibtex] [pdf] |
| [200] | Kaposi's Sarcoma-Associated Herpesvirus Viral Interferon Regulatory Factor 4 (vIRF4) Perturbs the G1-S Cell Cycle Progression via Deregulation of the cyclin D1 Gene. (Lee, H. R., Mitra, J., Lee, S., Gao, S. J., Oh, T. K., Kim, M. H., Ha, T. and Jung, J. U.), In J. Virol., 2016. [bibtex] [pdf] |
| 2015 | |
| [199] | Effects of DNA replication on mRNA noise. (Peterson, J. R., Cole, J. A., Fei, J., Ha, T. and Luthey-Schulten, Z. A.), In Proc. Natl. Acad. Sci. U.S.A., 2015. [bibtex] [pdf] |
| [198] | Integrin Molecular Tension within Motile Focal Adhesions. (Wang, X., Sun, J., Xu, Q., Chowdhury, F., Roein-Peikar, M., Wang, Y. and Ha, T.), In Biophys. J., 2015. [bibtex] [pdf] |
| [197] | Tandem Spinach Array for mRNA Imaging in Living Bacterial Cells. (Zhang, J., Fei, J., Leslie, B. J., Han, K. Y., Kuhlman, T. E. and Ha, T.), In Sci Rep, 2015. [bibtex] [pdf] |
| [196] | Single molecular force across single integrins dictates cell spreading. (Chowdhury, F., Li, I. T. S., Leslie, B. J., Doğanay, S., Singh, R., Wang, X., Seong, J., Lee, S. H., Park, S., Wang, N. and Ha, T.), In Integr Biol (Camb), 2015. [bibtex] [pdf] |
| [195] | Allosteric Regulation of E-Cadherin Adhesion. (Shashikanth, N., Petrova, Y. I., Park, S., Chekan, J., Maiden, S., Spano, M., Ha, T., Gumbiner, B. M. and Leckband, D. E.), In J. Biol. Chem., 2015. [bibtex] [pdf] |
| [194] | BEND3 represses rDNA transcription by stabilizing a NoRC component via USP21 deubiquitinase. (Khan, A., Giri, S., Wang, Y., Chakraborty, A., Ghosh, A. K., Anantharaman, A., Aggarwal, V., Sathyan, K. M., Ha, T., Prasanth, K. V. and Prasanth, S. G.), In Proc. Natl. Acad. Sci. U.S.A., 2015. [bibtex] [pdf] |
| [193] | A parameter estimation method for fluorescence lifetime data. (Sewell, D., Kim, H., Ha, T. and Ma, P.), In BMC Res Notes, 2015. [bibtex] [pdf] |
| [192] | Dual-color three-dimensional STED microscopy with a single high-repetition-rate laser. (Han, K. Y. and Ha, T.), In Opt Lett, 2015. [bibtex] [pdf] |
| [191] | Nucleosomes undergo slow spontaneous gaping. (Ngo, T. T. and Ha, T.), In Nucleic Acids Res., 2015. [bibtex] [pdf] |
| [190] | The preRC protein ORCA organizes heterochromatin by assembling histone H3 lysine 9 methyltransferases on chromatin. (Giri, S., Aggarwal, V., Pontis, J., Shen, Z., Chakraborty, A., Khan, A., Mizzen, C., Prasanth, K. V., Ait-Si-Ali, S., Ha, T. and Prasanth, S. G.), In Elife, 2015. [bibtex] [pdf] |
| [189] | Protein structure. Direct observation of structure-function relationship in a nucleic acid-processing enzyme. (Comstock, M. J., Whitley, K. D., Jia, H., Sokoloski, J., Lohman, T. M., Ha, T. and Chemla, Y. R.), In Science, 2015. [bibtex] [pdf] |
| [188] | Protein structure. Engineering of a superhelicase through conformational control. (Arslan, S., Khafizov, R., Thomas, C. D., Chemla, Y. R. and Ha, T.), In Science, 2015. [bibtex] [pdf] |
| [187] | RNA biochemistry. Determination of in vivo target search kinetics of regulatory noncoding RNA. (Fei, J., Singh, D., Zhang, Q., Park, S., Balasubramanian, D., Golding, I., Vanderpool, C. K. and Ha, T.), In Science, 2015. [bibtex] [pdf] |
| [186] | Asymmetric unwrapping of nucleosomes under tension directed by DNA local flexibility. (Ngo, T. T., Zhang, Q., Zhou, R., Yodh, J. G. and Ha, T.), In Cell, 2015. [bibtex] [pdf] |
| [185] | Distinct mechanisms regulating mechanical force-induced Ca²⁺ signals at the plasma membrane and the ER in human MSCs. (Kim, T. J., Joo, C., Seong, J., Vafabakhsh, R., Botvinick, E. L., Berns, M. W., Palmer, A. E., Wang, N., Ha, T., Jakobsson, E., Sun, J. and Wang, Y.), In Elife, 2015. [bibtex] [pdf] |
| [184] | Dynamic growth and shrinkage govern the pH dependence of RecA filament stability. (Kim, S. H., Park, J., Joo, C., Kim, D. and Ha, T.), In PLoS ONE, 2015. [bibtex] [pdf] |
| [183] | RNA fluorescence in situ hybridization in cultured mammalian cells. (Tripathi, V., Fei, J., Ha, T. and Prasanth, K. V.), In Methods Mol. Biol., 2015. [bibtex] [pdf] |
| 2014 | |
| [182] | A price to pay for relaxed substrate specificity: a comparative kinetic analysis of the class II lanthipeptide synthetases ProcM and HalM2. (Thibodeaux, C. J., Ha, T. and van der Donk, W. A.), In J. Am. Chem. Soc., 2014. [bibtex] [pdf] |
| [181] | Stoichiometry and assembly of mTOR complexes revealed by single-molecule pulldown. (Jain, A., Arauz, E., Aggarwal, V., Ikon, N., Chen, J. and Ha, T.), In Proc. Natl. Acad. Sci. U.S.A., 2014. [bibtex] [pdf] |
| [180] | An improved surface passivation method for single-molecule studies. (Hua, B., Han, K. Y., Zhou, R., Kim, H., Shi, X., Abeysirigunawardena, S. C., Jain, A., Singh, D., Aggarwal, V., Woodson, S. A. and Ha, T.), In Nat. Methods, 2014. [bibtex] [pdf] |
| [179] | Targeting DNA G-quadruplexes with helical small molecules. (Müller, S., Laxmi-Reddy, K., Jena, P. V., Baptiste, B., Dong, Z., Godde, F., Ha, T., Rodriguez, R., Balasubramanian, S. and Huc, I.), In Chembiochem, 2014. [bibtex] [pdf] |
| [178] | Single-molecule pull-down (SiMPull) for new-age biochemistry: methodology and biochemical applications of single-molecule pull-down (SiMPull) for probing biomolecular interactions in crude cell extracts. (Aggarwal, V. and Ha, T.), In Bioessays, 2014. [bibtex] [pdf] |
| [177] | Designing a nine cysteine-less DNA packaging motor from bacteriophage T4 reveals new insights into ATPase structure and function. (Kondabagil, K., Dai, L., Vafabakhsh, R., Ha, T., Draper, B. and Rao, V. B.), In Virology, 2014. [bibtex] [pdf] |
| [176] | Cooperative conformational transitions keep RecA filament active during ATPase cycle. (Kim, S. H., Ragunathan, K., Park, J., Joo, C., Kim, D. and Ha, T.), In J. Am. Chem. Soc., 2014. [bibtex] [pdf] |
| [175] | Single-molecule packaging initiation in real time by a viral DNA packaging machine from bacteriophage T4. (Vafabakhsh, R., Kondabagil, K., Earnest, T., Lee, K. S., Zhang, Z., Dai, L., Dahmen, K. A., Rao, V. B. and Ha, T.), In Proc. Natl. Acad. Sci. U.S.A., 2014. [bibtex] [pdf] |
| [174] | Single-molecule methods leap ahead. (Ha, T.), In Nat. Methods, 2014. [bibtex] [pdf] |
| [173] | A Coarse-Grained Model of Unstructured Single-Stranded DNA Derived from Atomistic Simulation and Single-Molecule Experiment. (Maffeo, C., Ngo, T. T., Ha, T. and Aksimentiev, A.), In J Chem Theory Comput, 2014. [bibtex] [pdf] |
| [172] | Matrix softness regulates plasticity of tumour-repopulating cells via H3K9 demethylation and Sox2 expression. (Tan, Y., Tajik, A., Chen, J., Jia, Q., Chowdhury, F., Wang, L., Chen, J., Zhang, S., Hong, Y., Yi, H., Wu, D. C., Zhang, Y., Wei, F., Poh, Y. C., Seong, J., Singh, R., Lin, L. J., Doğanay, S., Li, Y., Jia, H., Ha, T., Wang, Y., Huang, B. and Wang, N.), In Nat Commun, 2014. [bibtex] [pdf] |
| [171] | Biophysics. Ultraslow relaxation of confined DNA. (Chemla, Y. R. and Ha, T.), In Science, 2014. [bibtex] [pdf] |
| [170] | Ultrafast redistribution of E. coli SSB along long single-stranded DNA via intersegment transfer. (Lee, K. S., Marciel, A. B., Kozlov, A. G., Schroeder, C. M., Lohman, T. M. and Ha, T.), In J. Mol. Biol., 2014. [bibtex] [pdf] |
| [169] | Periodic DNA patrolling underlies diverse functions of Pif1 on R-loops and G-rich DNA. (Zhou, R., Zhang, J., Bochman, M. L., Zakian, V. A. and Ha, T.), In Elife, 2014. [bibtex] [pdf] |
| [168] | Single molecule analysis of Thermus thermophilus SSB protein dynamics on single-stranded DNA. (Zhang, J., Zhou, R., Inoue, J., Mikawa, T. and Ha, T.), In Nucleic Acids Res., 2014. [bibtex] [pdf] |
| [167] | Single-molecule fluorescence reveals the unwinding stepping mechanism of replicative helicase. (Syed, S., Pandey, M., Patel, S. S. and Ha, T.), In Cell Rep, 2014. [bibtex] [pdf] |
| [166] | Dynamic look at DNA unwinding by a replicative helicase. (Lee, S. J., Syed, S., Enemark, E. J., Schuck, S., Stenlund, A., Ha, T. and Joshua-Tor, L.), In Proc. Natl. Acad. Sci. U.S.A., 2014. [bibtex] [pdf] |
| [165] | Single-molecule optical spectroscopy. (Orrit, M., Ha, T. and Sandoghdar, V.), In Chem Soc Rev, 2014. [bibtex] [pdf] |
| [164] | Protein-guided RNA dynamics during early ribosome assembly. (Kim, H., Abeysirigunawarden, S. C., Chen, K., Mayerle, M., Ragunathan, K., Luthey-Schulten, Z., Ha, T. and Woodson, S. A.), In Nature, 2014. [bibtex] [pdf] |
| [163] | Crosstalk between the cGAS DNA sensor and Beclin-1 autophagy protein shapes innate antimicrobial immune responses. (Liang, Q., Seo, G. J., Choi, Y. J., Kwak, M. J., Ge, J., Rodgers, M. A., Shi, M., Leslie, B. J., Hopfner, K. P., Ha, T., Oh, B. H. and Jung, J. U.), In Cell Host Microbe, 2014. [bibtex] [pdf] |
| [162] | Kaposi's sarcoma-associated herpesvirus viral interferon regulatory factor 4 (vIRF4) targets expression of cellular IRF4 and the Myc gene to facilitate lytic replication. (Lee, H. R., Doğanay, S., Chung, B., Toth, Z., Brulois, K., Lee, S., Kanketayeva, Z., Feng, P., Ha, T. and Jung, J. U.), In J. Virol., 2014. [bibtex] [pdf] |
| [161] | Structural mechanisms of PriA-mediated DNA replication restart. (Bhattacharyya, B., George, N. P., Thurmes, T. M., Zhou, R., Jani, N., Wessel, S. R., Sandler, S. J., Ha, T. and Keck, J. L.), In Proc. Natl. Acad. Sci. U.S.A., 2014. [bibtex] [pdf] |
| [160] | Relaxed rotational and scrunching changes in P266L mutant of T7 RNA polymerase reduce short abortive RNAs while delaying transition into elongation. (Tang, G. Q., Nandakumar, D., Bandwar, R. P., Lee, K. S., Roy, R., Ha, T. and Patel, S. S.), In PLoS ONE, 2014. [bibtex] [pdf] |
| [159] | Towards a computational model of a methane producing archaeum. (Peterson, J. R., Labhsetwar, P., Ellermeier, J. R., Kohler, P. R., Jain, A., Ha, T., Metcalf, W. W. and Luthey-Schulten, Z.), In Archaea, 2014. [bibtex] [pdf] |
| 2013 | |
| [158] | Understanding the photophysics of the spinach-DFHBI RNA aptamer-fluorogen complex to improve live-cell RNA imaging. (Han, K. Y., Leslie, B. J., Fei, J., Zhang, J. and Ha, T.), In J. Am. Chem. Soc., 2013. [bibtex] [pdf] |
| [157] | Watching DNA breath one molecule at a time. (Fei, J. and Ha, T.), In Proc. Natl. Acad. Sci. U.S.A., 2013. [bibtex] [pdf] |
| [156] | ATPase-driven oligomerization of RIG-I on RNA allows optimal activation of type-I interferon. (Patel, J. R., Jain, A., Chou, Y. Y., Baum, A., Ha, T. and García-Sastre, A.), In EMBO Rep., 2013. [bibtex] [pdf] |
| [155] | Molecular mechanism of sequence-dependent stability of RecA filament. (Kim, S. H., Joo, C., Ha, T. and Kim, D.), In Nucleic Acids Res., 2013. [bibtex] [pdf] |
| [154] | Single-molecule approaches embrace molecular cohorts. (Ha, T.), In Cell, 2013. [bibtex] [pdf] |
| [153] | A dendritic single-molecule fluorescent probe that is monovalent, photostable and minimally blinking. (Yang, S. K., Shi, X., Park, S., Ha, T. and Zimmerman, S. C.), In Nat Chem, 2013. [bibtex] [pdf] |
| [152] | Single-molecule FRET reveals the native-state dynamics of the IκBα ankyrin repeat domain. (Lamboy, J. A., Kim, H., Dembinski, H., Ha, T. and Komives, E. A.), In J. Mol. Biol., 2013. [bibtex] [pdf] |
| [151] | PriC-mediated DNA replication restart requires PriC complex formation with the single-stranded DNA-binding protein. (Wessel, S. R., Marceau, A. H., Massoni, S. C., Zhou, R., Ha, T., Sandler, S. J. and Keck, J. L.), In J. Biol. Chem., 2013. [bibtex] [pdf] |
| [150] | Defining single molecular forces required to activate integrin and notch signaling. (Wang, X. and Ha, T.), In Science, 2013. [bibtex] [pdf] |
| [149] | Antibiotics that bind to the A site of the large ribosomal subunit can induce mRNA translocation. (Ermolenko, D. N., Cornish, P. V., Ha, T. and Noller, H. F.), In RNA, 2013. [bibtex] [pdf] |
| [148] | Fusion pore formation and expansion induced by Ca2+ and synaptotagmin 1. (Lai, Y., Diao, J., Liu, Y., Ishitsuka, Y., Su, Z., Schulten, K., Ha, T. and Shin, Y. K.), In Proc. Natl. Acad. Sci. U.S.A., 2013. [bibtex] [pdf] |
| [147] | The SOSS1 single-stranded DNA binding complex promotes DNA end resection in concert with Exo1. (Yang, S. H., Zhou, R., Campbell, J., Chen, J., Ha, T. and Paull, T. T.), In EMBO J., 2013. [bibtex] [pdf] |
| [146] | Single-molecule nanometry for biological physics. (Kim, H. and Ha, T.), In Rep Prog Phys, 2013. [bibtex] [pdf] |
| [145] | Direct imaging of single UvrD helicase dynamics on long single-stranded DNA. (Lee, K. S., Balci, H., Jia, H., Lohman, T. M. and Ha, T.), In Nat Commun, 2013. [bibtex] [pdf] |
| [144] | The telomere capping complex CST has an unusual stoichiometry, makes multipartite interaction with G-Tails, and unfolds higher-order G-tail structures. (Lue, N. F., Zhou, R., Chico, L., Mao, N., Steinberg-Neifach, O. and Ha, T.), In PLoS Genet., 2013. [bibtex] [pdf] |
| 2012 | |
| [143] | RecA filament sliding on DNA facilitates homology search. (Ragunathan, K., Liu, C. and Ha, T.), In Elife, 2012. [bibtex] [pdf] |
| [142] | Activated GTPase movement on an RNA scaffold drives co-translational protein targeting. (Shen, K., Arslan, S., Akopian, D., Ha, T. and Shan, S. O.), In Nature, 2012. [bibtex] [pdf] |
| [141] | Single-molecule FRET with total internal reflection microscopy. (Joo, C. and Ha, T.), In Cold Spring Harb Protoc, 2012. [bibtex] [pdf] |
| [140] | Prism-type total internal reflection microscopy for single-molecule FRET. (Joo, C. and Ha, T.), In Cold Spring Harb Protoc, 2012. [bibtex] [pdf] |
| [139] | Objective-type total internal reflection microscopy (emission) for single-molecule FRET. (Joo, C. and Ha, T.), In Cold Spring Harb Protoc, 2012. [bibtex] [pdf] |
| [138] | Objective-type total internal reflection microscopy (excitation) for single-molecule FRET. (Joo, C. and Ha, T.), In Cold Spring Harb Protoc, 2012. [bibtex] [pdf] |
| [137] | A helicase with an extra spring in its step. (Schlierf, M. and Ha, T.), In Cell, 2012. [bibtex] [pdf] |
| [136] | Imaging and identifying impurities in single-molecule FRET studies. (Joo, C. and Ha, T.), In Cold Spring Harb Protoc, 2012. [bibtex] [pdf] |
| [135] | Preparing sample chambers for single-molecule FRET. (Joo, C. and Ha, T.), In Cold Spring Harb Protoc, 2012. [bibtex] [pdf] |
| [134] | Dynamics of major histocompatibility complex class I association with the human peptide-loading complex. (Panter, M. S., Jain, A., Leonhardt, R. M., Ha, T. and Cresswell, P.), In J. Biol. Chem., 2012. [bibtex] [pdf] |
| [133] | Labeling proteins for single-molecule FRET. (Joo, C. and Ha, T.), In Cold Spring Harb Protoc, 2012. [bibtex] [pdf] |
| [132] | Labeling DNA (or RNA) for single-molecule FRET. (Joo, C. and Ha, T.), In Cold Spring Harb Protoc, 2012. [bibtex] [pdf] |
| [131] | Extreme bendability of DNA less than 100 base pairs long revealed by single-molecule cyclization. (Vafabakhsh, R. and Ha, T.), In Science, 2012. [bibtex] [pdf] |
| [130] | Dynamic association of ORCA with prereplicative complex components regulates DNA replication initiation. (Shen, Z., Chakraborty, A., Jain, A., Giri, S., Ha, T., Prasanth, K. V. and Prasanth, S. G.), In Mol. Cell. Biol., 2012. [bibtex] [pdf] |
| [129] | The influenza A virus PB2, PA, NP, and M segments play a pivotal role during genome packaging. (Gao, Q., Chou, Y. Y., Doğanay, S., Vafabakhsh, R., Ha, T. and Palese, P.), In J. Virol., 2012. [bibtex] [pdf] |
| [128] | Elastic coupling between RNA degradation and unwinding by an exoribonuclease. (Lee, G., Bratkowski, M. A., Ding, F., Ke, A. and Ha, T.), In Science, 2012. [bibtex] [pdf] |
| [127] | Single-molecule imaging: a collagenase pauses before embarking on a killing spree. (Lee, G. and Ha, T.), In Curr. Biol., 2012. [bibtex] [pdf] |
| [126] | Assembly of the five-way junction in the ribosomal small subunit using hybrid MD-Gō simulations. (Chen, K., Eargle, J., Lai, J., Kim, H., Abeysirigunawardena, S., Mayerle, M., Woodson, S., Ha, T. and Luthey-Schulten, Z.), In J Phys Chem B, 2012. [bibtex] [pdf] |
| [125] | One influenza virus particle packages eight unique viral RNAs as shown by FISH analysis. (Chou, Y. Y., Vafabakhsh, R., Doğanay, S., Gao, Q., Ha, T. and Palese, P.), In Proc. Natl. Acad. Sci. U.S.A., 2012. [bibtex] [pdf] |
| [124] | A rule of seven in Watson-Crick base-pairing of mismatched sequences. (Cisse, I. I., Kim, H. and Ha, T.), In Nat. Struct. Mol. Biol., 2012. [bibtex] [pdf] |
| [123] | A single vesicle-vesicle fusion assay for in vitro studies of SNAREs and accessory proteins. (Diao, J., Ishitsuka, Y., Lee, H., Joo, C., Su, Z., Syed, S., Shin, Y. K., Yoon, T. Y. and Ha, T.), In Nat Protoc, 2012. [bibtex] [pdf] |
| [122] | Quantitative fluorescence labeling of aldehyde-tagged proteins for single-molecule imaging. (Shi, X., Jung, Y., Lin, L. J., Liu, C., Wu, C., Cann, I. K. and Ha, T.), In Nat. Methods, 2012. [bibtex] [pdf] |
| [121] | Single-molecule pull-down for studying protein interactions. (Jain, A., Liu, R., Xiang, Y. K. and Ha, T.), In Nat Protoc, 2012. [bibtex] [pdf] |
| [120] | Opening-closing dynamics of the mitochondrial transcription pre-initiation complex. (Kim, H., Tang, G. Q., Patel, S. S. and Ha, T.), In Nucleic Acids Res., 2012. [bibtex] [pdf] |
| [119] | Single-molecule views of protein movement on single-stranded DNA. (Ha, T., Kozlov, A. G. and Lohman, T. M.), In Annu Rev Biophys, 2012. [bibtex] [pdf] |
| [118] | Photophysics of fluorescent probes for single-molecule biophysics and super-resolution imaging. (Ha, T. and Tinnefeld, P.), In Annu Rev Phys Chem, 2012. [bibtex] [pdf] |
| [117] | Single-molecule analysis of SSB dynamics on single-stranded DNA. (Zhou, R. and Ha, T.), In Methods Mol. Biol., 2012. [bibtex] [pdf] |
| 2011 | |
| [116] | Antiparallel EmrE exports drugs by exchanging between asymmetric structures. (Morrison, E. A., DeKoster, G. T., Dutta, S., Vafabakhsh, R., Clarkson, M. W., Bahl, A., Kern, D., Ha, T. and Henzler-Wildman, K. A.), In Nature, 2011. [bibtex] [pdf] |
| [115] | Detecting intramolecular conformational dynamics of single molecules in short distance range with subnanometer sensitivity. (Zhou, R., Kunzelmann, S., Webb, M. R. and Ha, T.), In Nano Lett., 2011. [bibtex] [pdf] |
| [114] | An entirely specific type I A-kinase anchoring protein that can sequester two molecules of protein kinase A at mitochondria. (Means, C. K., Lygren, B., Langeberg, L. K., Jain, A., Dixon, R. E., Vega, A. L., Gold, M. G., Petrosyan, S., Taylor, S. S., Murphy, A. N., Ha, T., Santana, L. F., Tasken, K. and Scott, J. D.), In Proc. Natl. Acad. Sci. U.S.A., 2011. [bibtex] [pdf] |
| [113] | A single-molecule view of chaperonin cooperativity. (Ha, T. and Myong, S.), In Proc. Natl. Acad. Sci. U.S.A., 2011. [bibtex] [pdf] |
| [112] | Rotations of the 2B sub-domain of E. coli UvrD helicase/translocase coupled to nucleotide and DNA binding. (Jia, H., Korolev, S., Niedziela-Majka, A., Maluf, N. K., Gauss, G. H., Myong, S., Ha, T., Waksman, G. and Lohman, T. M.), In J. Mol. Biol., 2011. [bibtex] [pdf] |
| [111] | Single-molecule nanopositioning: structural transitions of a helicase-DNA complex during ATP hydrolysis. (Balci, H., Arslan, S., Myong, S., Lohman, T. M. and Ha, T.), In Biophys. J., 2011. [bibtex] [pdf] |
| [110] | Real-time observation of strand exchange reaction with high spatiotemporal resolution. (Ragunathan, K., Joo, C. and Ha, T.), In Structure, 2011. [bibtex] [pdf] |
| [109] | Tyrosine phosphorylation enhances RAD52-mediated annealing by modulating its DNA binding. (Honda, M., Okuno, Y., Yoo, J., Ha, T. and Spies, M.), In EMBO J., 2011. [bibtex] [pdf] |
| [108] | SSB functions as a sliding platform that migrates on DNA via reptation. (Zhou, R., Kozlov, A. G., Roy, R., Zhang, J., Korolev, S., Lohman, T. M. and Ha, T.), In Cell, 2011. [bibtex] [pdf] |
| [107] | Monovalent, clickable, uncharged, water-soluble perylenediimide-cored dendrimers for target-specific fluorescent biolabeling. (Yang, S. K., Shi, X., Park, S., Doganay, S., Ha, T. and Zimmerman, S. C.), In J. Am. Chem. Soc., 2011. [bibtex] [pdf] |
| [106] | Visualization of the nanospring dynamics of the IkappaBalpha ankyrin repeat domain in real time. (Lamboy, J. A., Kim, H., Lee, K. S., Ha, T. and Komives, E. A.), In Proc. Natl. Acad. Sci. U.S.A., 2011. [bibtex] [pdf] |
| [105] | Editorial on nanobiology. (Kim, S. K., Ha, T. and Schermann, J. P.), In Phys Chem Chem Phys, 2011. [bibtex] [pdf] |
| [104] | Single-molecule analysis reveals three phases of DNA degradation by an exonuclease. (Lee, G., Yoo, J., Leslie, B. J. and Ha, T.), In Nat. Chem. Biol., 2011. [bibtex] [pdf] |
| [103] | Probing cellular protein complexes using single-molecule pull-down. (Jain, A., Liu, R., Ramani, B., Arauz, E., Ishitsuka, Y., Ragunathan, K., Park, J., Chen, J., Xiang, Y. K. and Ha, T.), In Nature, 2011. [bibtex] [pdf] |
| [102] | Forcing a connection: impacts of single-molecule force spectroscopy on in vivo tension sensing. (Brenner, M. D., Zhou, R. and Ha, T.), In Biopolymers, 2011. [bibtex] [pdf] |
| [101] | Ultrahigh-resolution optical trap with single-fluorophore sensitivity. (Comstock, M. J., Ha, T. and Chemla, Y. R.), In Nat. Methods, 2011. [bibtex] [pdf] |
| [100] | Reverse-chaperoning activity of an AAA+ protein. (Liu, C., McKinney, M. C., Chen, Y. H., Earnest, T. M., Shi, X., Lin, L. J., Ishino, Y., Dahmen, K., Cann, I. K. and Ha, T.), In Biophys. J., 2011. [bibtex] [pdf] |
| [99] | Seeing a molecular machine self-renew. (Shi, X. and Ha, T.), In Proc. Natl. Acad. Sci. U.S.A., 2011. [bibtex] [pdf] |
| [98] | A promiscuous DNA packaging machine from bacteriophage T4. (Zhang, Z., Kottadiel, V. I., Vafabakhsh, R., Dai, L., Chemla, Y. R., Ha, T. and Rao, V. B.), In PLoS Biol., 2011. [bibtex] [pdf] |
| [97] | Homochirality and the origin of life. Editorial of the PCCP themed issue. (Kim, S. K., Ha, T. and Schermann, J. P.), In Phys Chem Chem Phys, 2011. [bibtex] [pdf] |
| 2010 | |
| [96] | Single-molecule four-color FRET. (Lee, J., Lee, S., Ragunathan, K., Joo, C., Ha, T. and Hohng, S.), In Angew. Chem. Int. Ed. Engl., 2010. [bibtex] [pdf] |
| [95] | Reversible and controllable nanolocomotion of an RNA-processing machinery. (Lee, G., Hartung, S., Hopfner, K. P. and Ha, T.), In Nano Lett., 2010. [bibtex] [pdf] |
| [94] | Temperature-independent porous nanocontainers for single-molecule fluorescence studies. (Ishitsuka, Y., Okumus, B., Arslan, S., Chen, K. H. and Ha, T.), In Anal. Chem., 2010. [bibtex] [pdf] |
| [93] | 5'-Single-stranded/duplex DNA junctions are loading sites for E. coli UvrD translocase. (Tomko, E. J., Jia, H., Park, J., Maluf, N. K., Ha, T. and Lohman, T. M.), In EMBO J., 2010. [bibtex] [pdf] |
| [92] | Advances in mass spectrometry for biological science. (Kim, S. K., Ha, T. and Schermann, J. P.), In Phys Chem Chem Phys, 2010. [bibtex] [pdf] |
| [91] | The role of L1 stalk-tRNA interaction in the ribosome elongation cycle. (Trabuco, L. G., Schreiner, E., Eargle, J., Cornish, P., Ha, T., Luthey-Schulten, Z. and Schulten, K.), In J. Mol. Biol., 2010. [bibtex] [pdf] |
| [90] | Water in biological systems. (Kim, S. K., Ha, T. and Schermann, J. P.), In Phys Chem Chem Phys, 2010. [bibtex] [pdf] |
| [89] | PcrA helicase dismantles RecA filaments by reeling in DNA in uniform steps. (Park, J., Myong, S., Niedziela-Majka, A., Lee, K. S., Yu, J., Lohman, T. M. and Ha, T.), In Cell, 2010. [bibtex] [pdf] |
| [88] | A single-vesicle content mixing assay for SNARE-mediated membrane fusion. (Diao, J., Su, Z., Ishitsuka, Y., Lu, B., Lee, K. S., Lai, Y., Shin, Y. K. and Ha, T.), In Nat Commun, 2010. [bibtex] [pdf] |
| [87] | Acidification of the oxygen scavenging system in single-molecule fluorescence studies: in situ sensing with a ratiometric dual-emission probe. (Shi, X., Lim, J. and Ha, T.), In Anal. Chem., 2010. [bibtex] [pdf] |
| [86] | Measuring mechanical tension across vinculin reveals regulation of focal adhesion dynamics. (Grashoff, C., Hoffman, B. D., Brenner, M. D., Zhou, R., Parsons, M., Yang, M. T., McLean, M. A., Sligar, S. G., Chen, C. S., Ha, T. and Schwartz, M. A.), In Nature, 2010. [bibtex] [pdf] |
| [85] | Inching over hurdles: how DNA helicases move on crowded lattices. (Spies, M. and Ha, T.), In Cell Cycle, 2010. [bibtex] [pdf] |
| [84] | Human Rad52 binds and wraps single-stranded DNA and mediates annealing via two hRad52-ssDNA complexes. (Grimme, J. M., Honda, M., Wright, R., Okuno, Y., Rothenberg, E., Mazin, A. V., Ha, T. and Spies, M.), In Nucleic Acids Res., 2010. [bibtex] [pdf] |
| [83] | Insight into helicase mechanism and function revealed through single-molecule approaches. (Yodh, J. G., Schlierf, M. and Ha, T.), In Q. Rev. Biophys., 2010. [bibtex] [pdf] |
| [82] | A comparative study of multivariate and univariate hidden Markov modelings in time-binned single-molecule FRET data analysis. (Liu, Y., Park, J., Dahmen, K. A., Chemla, Y. R. and Ha, T.), In J Phys Chem B, 2010. [bibtex] [pdf] |
| [81] | Biomolecular structures: from isolated molecules to the cell crowded medium. (Kim, S. K., Ha, T. and Schermann, J. P.), In Phys Chem Chem Phys, 2010. [bibtex] [pdf] |
| [80] | Single-Vesicle Fusion Assay Reveals Munc18-1 Binding to the SNARE Core Is Sufficient for Stimulating Membrane Fusion. (Diao, J., Su, Z., Lu, X., Yoon, T. Y., Shin, Y. K. and Ha, T.), In ACS Chem Neurosci, 2010. [bibtex] [pdf] |
| [79] | Multivector fluorescence analysis of the xpt guanine riboswitch aptamer domain and the conformational role of guanine. (Brenner, M. D., Scanlan, M. S., Nahas, M. K., Ha, T. and Silverman, S. K.), In Biochemistry, 2010. [bibtex] [pdf] |
| [78] | Stepwise translocation of nucleic acid motors. (Myong, S. and Ha, T.), In Curr. Opin. Struct. Biol., 2010. [bibtex] [pdf] |
| [77] | Force-fluorescence spectroscopy at the single-molecule level. (Zhou, R., Schlierf, M. and Ha, T.), In Meth. Enzymol., 2010. [bibtex] [pdf] |
| [76] | Single-molecule FRET analysis of helicase functions. (Rothenberg, E. and Ha, T.), In Methods Mol. Biol., 2010. [bibtex] [pdf] |
| [75] | Real-time observation of G-quadruplex dynamics using single-molecule FRET microscopy. (Okumus, B. and Ha, T.), In Methods Mol. Biol., 2010. [bibtex] [pdf] |
| 2009 | |
| [74] | Real-time observation of the transition from transcription initiation to elongation of the RNA polymerase. (Tang, G. Q., Roy, R., Bandwar, R. P., Ha, T. and Patel, S. S.), In Proc. Natl. Acad. Sci. U.S.A., 2009. [bibtex] [pdf] |
| [73] | Coordinating DNA replication by means of priming loop and differential synthesis rate. (Pandey, M., Syed, S., Donmez, I., Patel, G., Ha, T. and Patel, S. S.), In Nature, 2009. [bibtex] [pdf] |
| [72] | SSB protein diffusion on single-stranded DNA stimulates RecA filament formation. (Roy, R., Kozlov, A. G., Lohman, T. M. and Ha, T.), In Nature, 2009. [bibtex] [pdf] |
| [71] | Single molecule nanocontainers made porous using a bacterial toxin. (Okumus, B., Arslan, S., Fengler, S. M., Myong, S. and Ha, T.), In J. Am. Chem. Soc., 2009. [bibtex] [pdf] |
| [70] | Single-molecule analysis reveals differential effect of ssDNA-binding proteins on DNA translocation by XPD helicase. (Honda, M., Park, J., Pugh, R. A., Ha, T. and Spies, M.), In Mol. Cell, 2009. [bibtex] [pdf] |
| [69] | G-quadruplex DNA bound by a synthetic ligand is highly dynamic. (Jena, P. V., Shirude, P. S., Okumus, B., Laxmi-Reddy, K., Godde, F., Huc, I., Balasubramanian, S. and Ha, T.), In J. Am. Chem. Soc., 2009. [bibtex] [pdf] |
| [68] | Fluorescent lifetime trajectories of a single fluorophore reveal reaction intermediates during transcription initiation. (Sorokina, M., Koh, H. R., Patel, S. S. and Ha, T.), In J. Am. Chem. Soc., 2009. [bibtex] [pdf] |
| [67] | C2AB: a molecular glue for lipid vesicles with a negatively charged surface. (Diao, J., Yoon, T. Y., Su, Z., Shin, Y. K. and Ha, T.), In Langmuir, 2009. [bibtex] [pdf] |
| [66] | Photonics: How nanocrystals lost their blink. (Ha, T.), In Nature, 2009. [bibtex] [pdf] |
| [65] | DNA nanotechnology: a nanomachine goes live. (Ishitsuka, Y. and Ha, T.), In Nat Nanotechnol, 2009. [bibtex] [pdf] |
| [64] | Following movement of the L1 stalk between three functional states in single ribosomes. (Cornish, P. V., Ermolenko, D. N., Staple, D. W., Hoang, L., Hickerson, R. P., Noller, H. F. and Ha, T.), In Proc. Natl. Acad. Sci. U.S.A., 2009. [bibtex] [pdf] |
| [63] | Cytosolic viral sensor RIG-I is a 5'-triphosphate-dependent translocase on double-stranded RNA. (Myong, S., Cui, S., Cornish, P. V., Kirchhofer, A., Gack, M. U., Jung, J. U., Hopfner, K. P. and Ha, T.), In Science, 2009. [bibtex] [pdf] |
| [62] | BLM helicase measures DNA unwound before switching strands and hRPA promotes unwinding reinitiation. (Yodh, J. G., Stevens, B. C., Kanagaraj, R., Janscak, P. and Ha, T.), In EMBO J., 2009. [bibtex] [pdf] |
| [61] | Multimodal optical sensing and analyte specificity using single-walled carbon nanotubes. (Heller, D. A., Jin, H., Martinez, B. M., Patel, D., Miller, B. M., Yeung, T. K., Jena, P. V., Höbartner, C., Ha, T., Silverman, S. K. and Strano, M. S.), In Nat Nanotechnol, 2009. [bibtex] [pdf] |
| 2008 | |
| [60] | Human Rad52-mediated homology search and annealing occurs by continuous interactions between overlapping nucleoprotein complexes. (Rothenberg, E., Grimme, J. M., Spies, M. and Ha, T.), In Proc. Natl. Acad. Sci. U.S.A., 2008. [bibtex] [pdf] |
| [59] | Roles of RIG-I N-terminal tandem CARD and splice variant in TRIM25-mediated antiviral signal transduction. (Gack, M. U., Kirchhofer, A., Shin, Y. C., Inn, K. S., Liang, C., Cui, S., Myong, S., Ha, T., Hopfner, K. P. and Jung, J. U.), In Proc. Natl. Acad. Sci. U.S.A., 2008. [bibtex] [pdf] |
| [58] | Engineering of functional replication protein a homologs based on insights into the evolution of oligonucleotide/oligosaccharide-binding folds. (Lin, Y., Lin, L. J., Sriratana, P., Coleman, K., Ha, T., Spies, M. and Cann, I. K.), In J. Bacteriol., 2008. [bibtex] [pdf] |
| [57] | Orientation dependence in fluorescent energy transfer between Cy3 and Cy5 terminally attached to double-stranded nucleic acids. (Iqbal, A., Arslan, S., Okumus, B., Wilson, T. J., Giraud, G., Norman, D. G., Ha, T. and Lilley, D. M.), In Proc. Natl. Acad. Sci. U.S.A., 2008. [bibtex] [pdf] |
| [56] | Complexin and Ca2+ stimulate SNARE-mediated membrane fusion. (Yoon, T. Y., Lu, X., Diao, J., Lee, S. M., Ha, T. and Shin, Y. K.), In Nat. Struct. Mol. Biol., 2008. [bibtex] [pdf] |
| [55] | The SNARE complex from yeast is partially unstructured on the membrane. (Su, Z., Ishitsuka, Y., Ha, T. and Shin, Y. K.), In Structure, 2008. [bibtex] [pdf] |
| [54] | Branch migration enzyme as a Brownian ratchet. (Rasnik, I., Jeong, Y. J., McKinney, S. A., Rajagopal, V., Patel, S. S. and Ha, T.), In EMBO J., 2008. [bibtex] [pdf] |
| [53] | Spontaneous intersubunit rotation in single ribosomes. (Cornish, P. V., Ermolenko, D. N., Noller, H. F. and Ha, T.), In Mol. Cell, 2008. [bibtex] [pdf] |
| [52] | Transcription initiation in a single-subunit RNA polymerase proceeds through DNA scrunching and rotation of the N-terminal subdomains. (Tang, G. Q., Roy, R., Ha, T. and Patel, S. S.), In Mol. Cell, 2008. [bibtex] [pdf] |
| [51] | A practical guide to single-molecule FRET. (Roy, R., Hohng, S. and Ha, T.), In Nat. Methods, 2008. [bibtex] [pdf] |
| [50] | Advances in single-molecule fluorescence methods for molecular biology. (Joo, C., Balci, H., Ishitsuka, Y., Buranachai, C. and Ha, T.), In Annu. Rev. Biochem., 2008. [bibtex] [pdf] |
| 2007 | |
| [49] | How directional translocation is regulated in a DNA helicase motor. (Yu, J., Ha, T. and Schulten, K.), In Biophys. J., 2007. [bibtex] [pdf] |
| [48] | Dissecting metal ion-dependent folding and catalysis of a single DNAzyme. (Kim, H. K., Rasnik, I., Liu, J., Ha, T. and Lu, Y.), In Nat. Chem. Biol., 2007. [bibtex] [pdf] |
| [47] | MCM forked substrate specificity involves dynamic interaction with the 5'-tail. (Rothenberg, E., Trakselis, M. A., Bell, S. D. and Ha, T.), In J. Biol. Chem., 2007. [bibtex] [pdf] |
| [46] | Fluorescence-force spectroscopy maps two-dimensional reaction landscape of the holliday junction. (Hohng, S., Zhou, R., Nahas, M. K., Yu, J., Schulten, K., Lilley, D. M. and Ha, T.), In Science, 2007. [bibtex] [pdf] |
| [45] | Fueling protein DNA interactions inside porous nanocontainers. (Cisse, I., Okumus, B., Joo, C. and Ha, T.), In Proc. Natl. Acad. Sci. U.S.A., 2007. [bibtex] [pdf] |
| [44] | Spring-loaded mechanism of DNA unwinding by hepatitis C virus NS3 helicase. (Myong, S., Bruno, M. M., Pyle, A. M. and Ha, T.), In Science, 2007. [bibtex] [pdf] |
| [43] | Need for speed: mechanical regulation of a replicative helicase. (Ha, T.), In Cell, 2007. [bibtex] [pdf] |
| [42] | Dynamic structural rearrangements between DNA binding modes of E. coli SSB protein. (Roy, R., Kozlov, A. G., Lohman, T. M. and Ha, T.), In J. Mol. Biol., 2007. [bibtex] [pdf] |
| [41] | Single-molecule conformational analysis of G-quadruplex formation in the promoter DNA duplex of the proto-oncogene c-kit. (Shirude, P. S., Okumus, B., Ying, L., Ha, T. and Balasubramanian, S.), In J. Am. Chem. Soc., 2007. [bibtex] [pdf] |
| [40] | A survey of single-molecule techniques in chemical biology. (Cornish, P. V. and Ha, T.), In ACS Chem. Biol., 2007. [bibtex] [pdf] |
| [39] | Spectroscopic observation of RNA chaperone activities of Hfq in post-transcriptional regulation by a small non-coding RNA. (Arluison, V., Hohng, S., Roy, R., Pellegrini, O., Régnier, P. and Ha, T.), In Nucleic Acids Res., 2007. [bibtex] [pdf] |
| 2006 | |
| [38] | Multiple intermediates in SNARE-induced membrane fusion. (Yoon, T. Y., Okumus, B., Zhang, F., Shin, Y. K. and Ha, T.), In Proc. Natl. Acad. Sci. U.S.A., 2006. [bibtex] [pdf] |
| [37] | Nonblinking and long-lasting single-molecule fluorescence imaging. (Rasnik, I., McKinney, S. A. and Ha, T.), In Nat. Methods, 2006. [bibtex] [pdf] |
| [36] | Structure-based model of the stepping motor of PcrA helicase. (Yu, J., Ha, T. and Schulten, K.), In Biophys. J., 2006. [bibtex] [pdf] |
| [35] | Analysis of single-molecule FRET trajectories using hidden Markov modeling. (McKinney, S. A., Joo, C. and Ha, T.), In Biophys. J., 2006. [bibtex] [pdf] |
| [34] | Methanosarcina acetivorans flap endonuclease 1 activity is inhibited by a cognate single-stranded-DNA-binding protein. (Lin, Y., Guzman, C. E., McKinney, M. C., Nair, S. K., Ha, T. and Cann, I. K.), In J. Bacteriol., 2006. [bibtex] [pdf] |
| [33] | Real-time observation of RecA filament dynamics with single monomer resolution. (Joo, C., McKinney, S. A., Nakamura, M., Rasnik, I., Myong, S. and Ha, T.), In Cell, 2006. [bibtex] [pdf] |
| [32] | Defocused orientation and position imaging (DOPI) of myosin V. (Toprak, E., Enderlein, J., Syed, S., McKinney, S. A., Petschek, R. G., Ha, T., Goldman, Y. E. and Selvin, P. R.), In Proc. Natl. Acad. Sci. U.S.A., 2006. [bibtex] [pdf] |
| [31] | Bridging conformational dynamics and function using single-molecule spectroscopy. (Myong, S., Stevens, B. C. and Ha, T.), In Structure, 2006. [bibtex] [pdf] |
| [30] | Single molecule nanometronome. (Buranachai, C., McKinney, S. A. and Ha, T.), In Nano Lett., 2006. [bibtex] [pdf] |
| [29] | Unraveling helicase mechanisms one molecule at a time. (Rasnik, I., Myong, S. and Ha, T.), In Nucleic Acids Res., 2006. [bibtex] [pdf] |
| 2005 | |
| [28] | Extreme conformational diversity in human telomeric DNA. (Lee, J. Y., Okumus, B., Kim, D. S. and Ha, T.), In Proc. Natl. Acad. Sci. U.S.A., 2005. [bibtex] [pdf] |
| [27] | Repetitive shuttling of a motor protein on DNA. (Myong, S., Rasnik, I., Joo, C., Lohman, T. M. and Ha, T.), In Nature, 2005. [bibtex] [pdf] |
| [26] | Surfaces and orientations: much to FRET about? (Rasnik, I., McKinney, S. A. and Ha, T.), In Acc. Chem. Res., 2005. [bibtex] [pdf] |
| [25] | Interhead distance measurements in myosin VI via SHRImP support a simplified hand-over-hand model. (Balci, H., Ha, T., Sweeney, H. L. and Selvin, P. R.), In Biophys. J., 2005. [bibtex] [pdf] |
| [24] | Single-molecule quantum-dot fluorescence resonance energy transfer. (Hohng, S. and Ha, T.), In Chemphyschem, 2005. [bibtex] [pdf] |
| [23] | Observing spontaneous branch migration of Holliday junctions one step at a time. (McKinney, S. A., Freeman, A. D., Lilley, D. M. and Ha, T.), In Proc. Natl. Acad. Sci. U.S.A., 2005. [bibtex] [pdf] |
| [22] | The euryarchaeota, nature's medium for engineering of single-stranded DNA-binding proteins. (Robbins, J. B., McKinney, M. C., Guzman, C. E., Sriratana, B., Fitz-Gibbon, S., Ha, T. and Cann, I. K.), In J. Biol. Chem., 2005. [bibtex] [pdf] |
| [21] | Stereospecific effects determine the structure of a four-way DNA junction. (Liu, J., Déclais, A. C., McKinney, S. A., Ha, T., Norman, D. G. and Lilley, D. M.), In Chem. Biol., 2005. [bibtex] [pdf] |
| 2004 | |
| [20] | Observation of internal cleavage and ligation reactions of a ribozyme. (Nahas, M. K., Wilson, T. J., Hohng, S., Jarvie, K., Lilley, D. M. and Ha, T.), In Nat. Struct. Mol. Biol., 2004. [bibtex] [pdf] |
| [19] | Vesicle encapsulation studies reveal that single molecule ribozyme heterogeneities are intrinsic. (Okumus, B., Wilson, T. J., Lilley, D. M. and Ha, T.), In Biophys. J., 2004. [bibtex] [pdf] |
| [18] | Exploring rare conformational species and ionic effects in DNA Holliday junctions using single-molecule spectroscopy. (Joo, C., McKinney, S. A., Lilley, D. M. and Ha, T.), In J. Mol. Biol., 2004. [bibtex] [pdf] |
| [17] | Single-molecule three-color FRET. (Hohng, S., Joo, C. and Ha, T.), In Biophys. J., 2004. [bibtex] [pdf] |
| [16] | Single-molecule high-resolution imaging with photobleaching. (Gordon, M. P., Ha, T. and Selvin, P. R.), In Proc. Natl. Acad. Sci. U.S.A., 2004. [bibtex] [pdf] |
| [15] | Structural dynamics and processing of nucleic acids revealed by single-molecule spectroscopy. (Ha, T.), In Biochemistry, 2004. [bibtex] [pdf] |
| [14] | Probing single-stranded DNA conformational flexibility using fluorescence spectroscopy. (Murphy, M. C., Rasnik, I., Cheng, W., Lohman, T. M. and Ha, T.), In Biophys. J., 2004. [bibtex] [pdf] |
| [13] | Functional analysis of multiple single-stranded DNA-binding proteins from Methanosarcina acetivorans and their effects on DNA synthesis by DNA polymerase BI. (Robbins, J. B., Murphy, M. C., White, B. A., Mackie, R. I., Ha, T. and Cann, I. K.), In J. Biol. Chem., 2004. [bibtex] [pdf] |
| [12] | DNA-binding orientation and domain conformation of the E. coli rep helicase monomer bound to a partial duplex junction: single-molecule studies of fluorescently labeled enzymes. (Rasnik, I., Myong, S., Cheng, W., Lohman, T. M. and Ha, T.), In J. Mol. Biol., 2004. [bibtex] [pdf] |
| [11] | Near-complete suppression of quantum dot blinking in ambient conditions. (Hohng, S. and Ha, T.), In J. Am. Chem. Soc., 2004. [bibtex] [pdf] |
| [10] | Discrete and heterogeneous rotational dynamics of single membrane probe dyes in gel phase supported lipid bilayer. (Stevens, B. C. and Ha, T.), In J Chem Phys, 2004. [bibtex] [pdf] |
| [9] | Conformational flexibility of four-way junctions in RNA. (Hohng, S., Wilson, T. J., Tan, E., Clegg, R. M., Lilley, D. M. and Ha, T.), In J. Mol. Biol., 2004. [bibtex] [pdf] |
| [8] | Conformational model of the Holliday junction transition deduced from molecular dynamics simulations. (Yu, J., Ha, T. and Schulten, K.), In Nucleic Acids Res., 2004. [bibtex] [pdf] |
| 2003 | |
| [7] | A four-way junction accelerates hairpin ribozyme folding via a discrete intermediate. (Tan, E., Wilson, T. J., Nahas, M. K., Clegg, R. M., Lilley, D. M. and Ha, T.), In Proc. Natl. Acad. Sci. U.S.A., 2003. [bibtex] [pdf] |
| [6] | Myosin V walks hand-over-hand: single fluorophore imaging with 1.5-nm localization. (Yildiz, A., Forkey, J. N., McKinney, S. A., Ha, T., Goldman, Y. E. and Selvin, P. R.), In Science, 2003. [bibtex] [pdf] |
| [5] | Photodestruction intermediates probed by an adjacent reporter molecule. (Ha, T. and Xu, J.), In Phys. Rev. Lett., 2003. [bibtex] [pdf] |
| [4] | Structural dynamics of individual Holliday junctions. (McKinney, S. A., Déclais, A. C., Lilley, D. M. and Ha, T.), In Nat. Struct. Biol., 2003. [bibtex] [pdf] |
| 2002 | |
| [3] | Early steps of supported bilayer formation probed by single vesicle fluorescence assays. (Johnson, J. M., Ha, T., Chu, S. and Boxer, S. G.), In Biophys. J., 2002. [bibtex] [pdf] |
| [2] | Initiation and re-initiation of DNA unwinding by the Escherichia coli Rep helicase. (Ha, T., Rasnik, I., Cheng, W., Babcock, H. P., Gauss, G. H., Lohman, T. M. and Chu, S.), In Nature, 2002. [bibtex] [pdf] |
| [1] | Mg2+-dependent conformational change of RNA studied by fluorescence correlation and FRET on immobilized single molecules. (Kim, H. D., Nienhaus, G. U., Ha, T., Orr, J. W., Williamson, J. R. and Chu, S.), In Proc. Natl. Acad. Sci. U.S.A., 2002. [bibtex] [pdf] |